Pavan Ramkumar
Departments of Neurobiology and Physical Medicine & Rehabilitation, Northwestern University
Rehabilitation Institute of Chicago, 345 East Superior Street Chicago IL, 60611 Phone: (312) 608-7178
Unprecedented advances in both experimental techniques to monitor brain function, and computational infrastructure and algorithms over the past decade, provide tremendous opportunities to reverse engineer the brain basis of perception and behavior. To transition our efforts from a pre-Galilean age of individualized discoveries into an era of integrated theory development, we need to make two major computational advances. First, we need to develop models of perception and behavior that generate testable neurobiological predictions. Second, we need to analyze data from neuroscience experiments to test these predictions. I work at the intersection of these two computational endeavors.
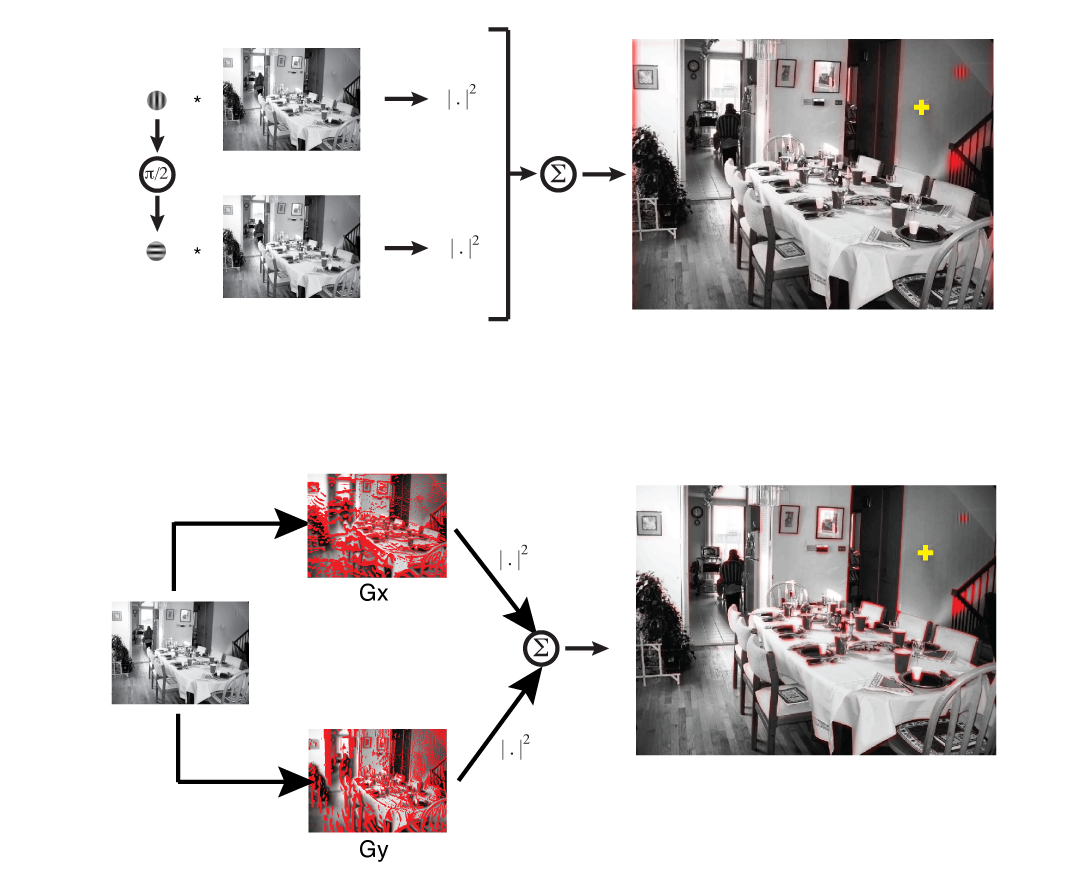
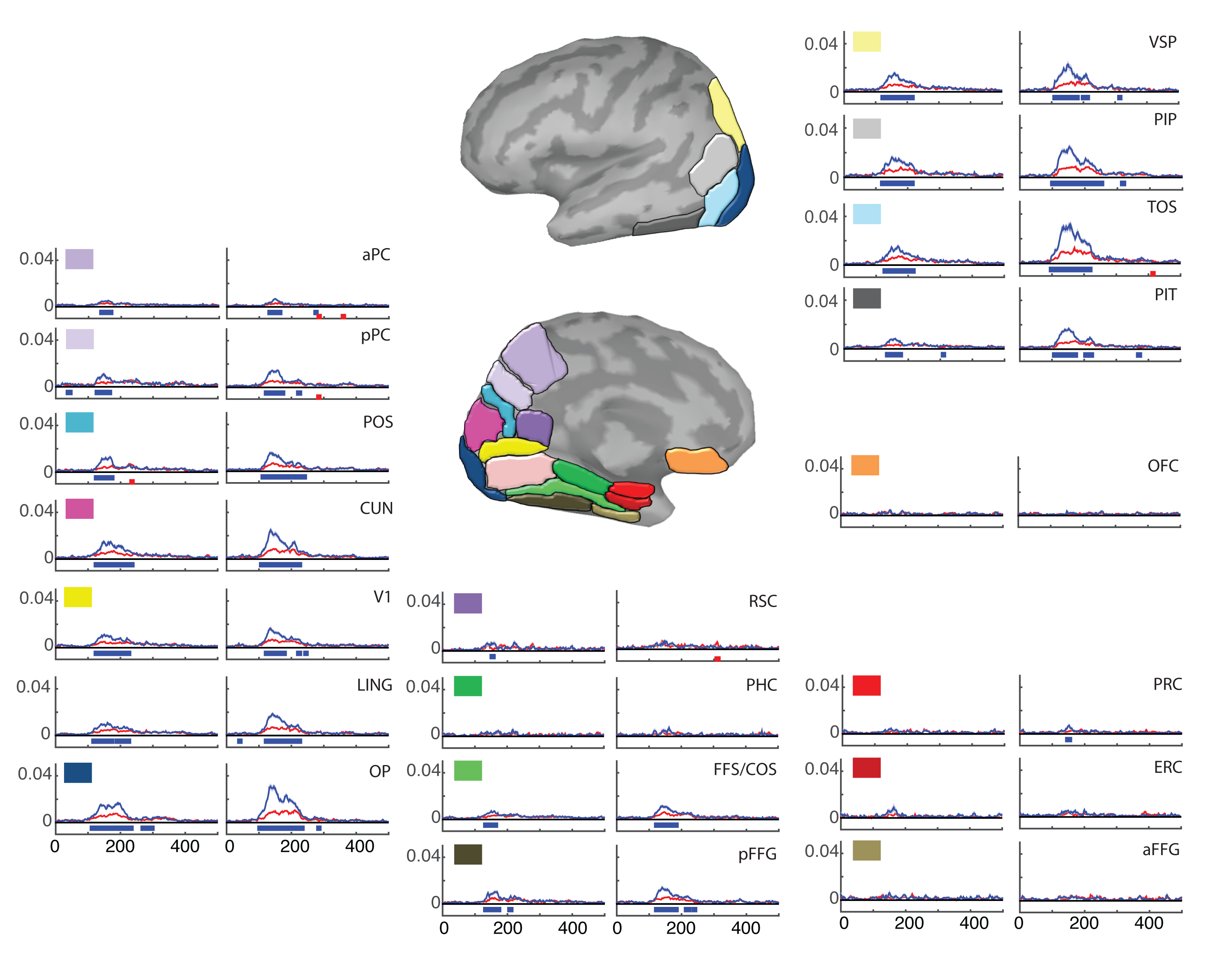
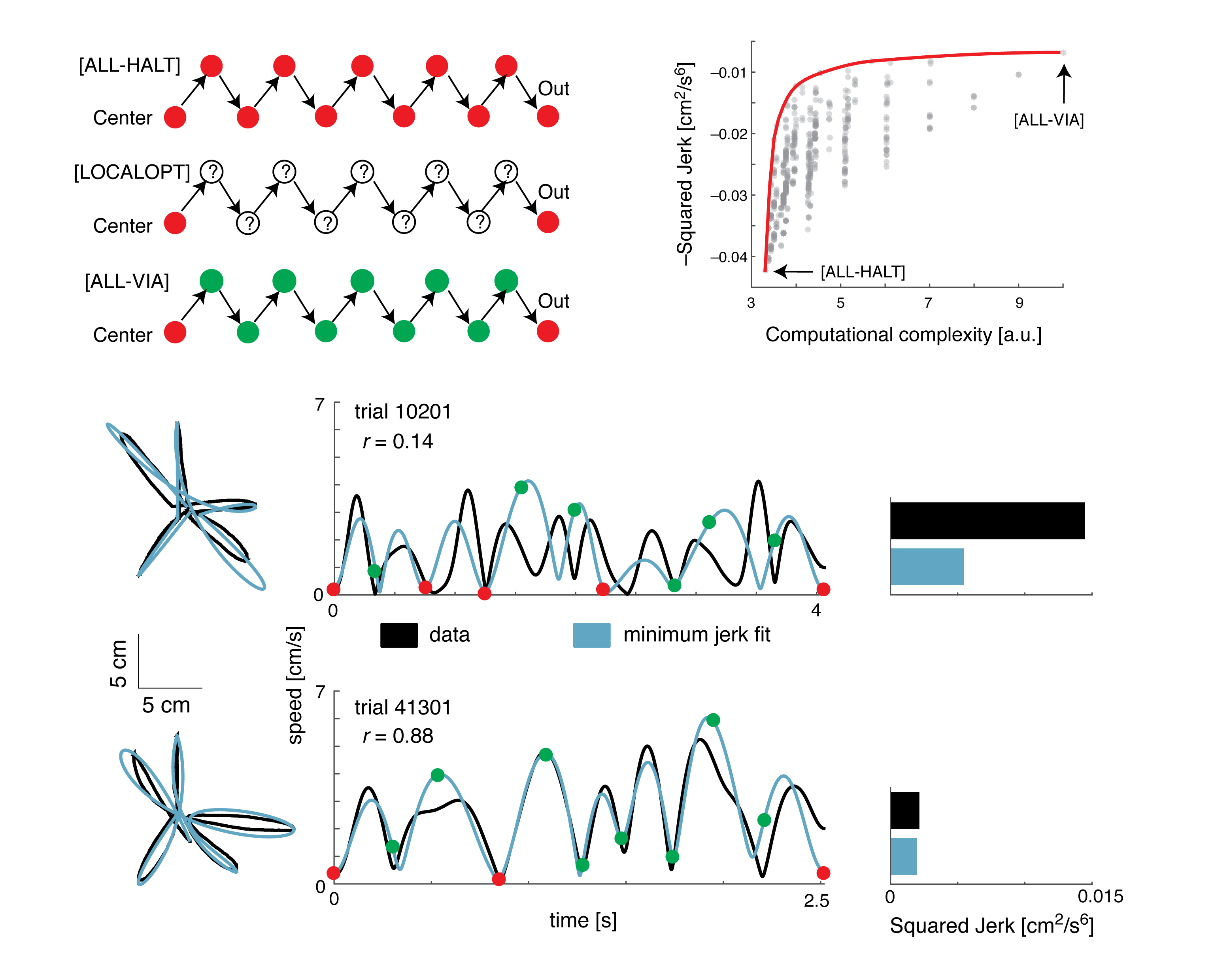
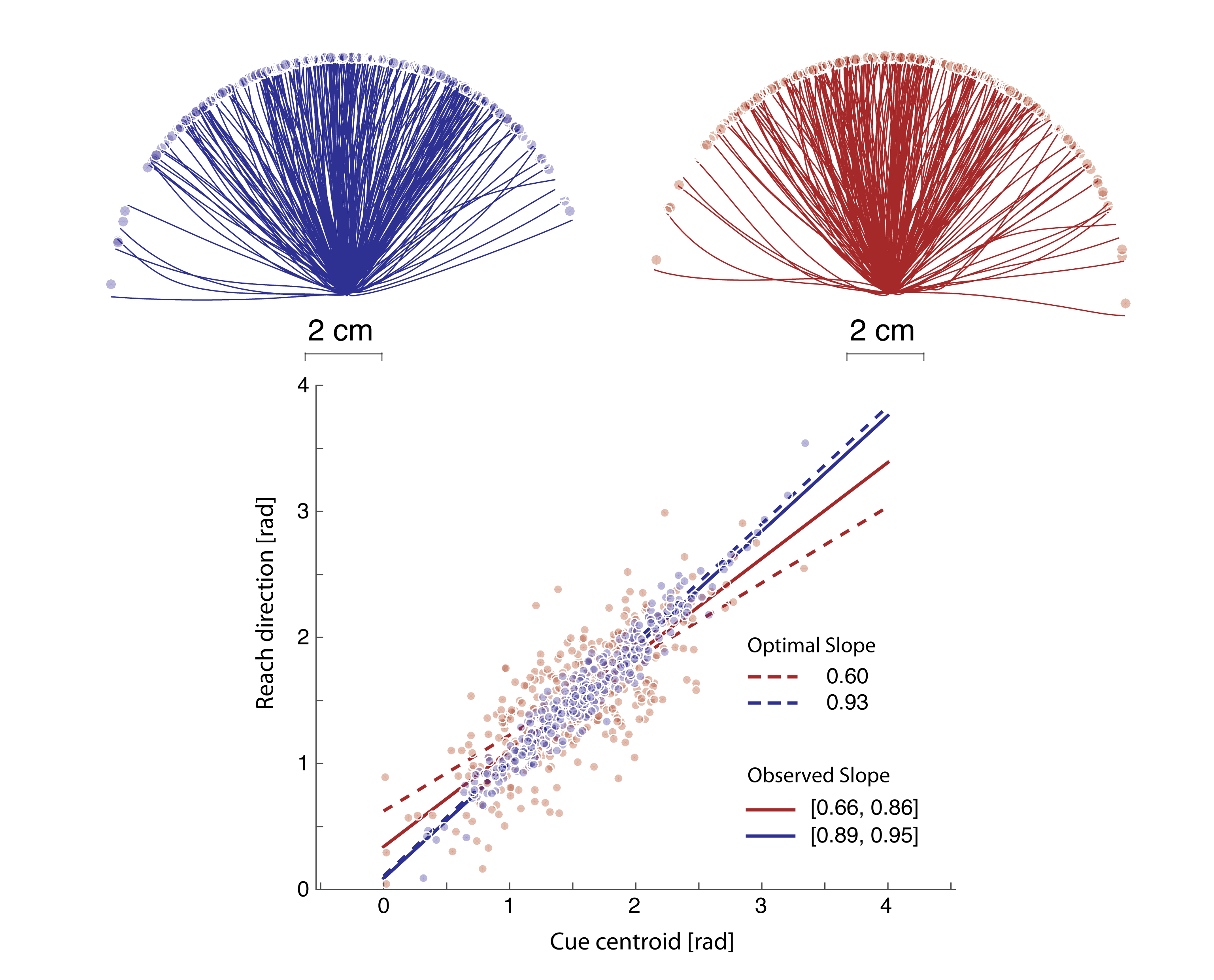
A computational lens into brain function enables us to formalize perception and behavior as the result of neural computations. This branch of my research brings computational motor control to the brain basis of movement and computer vision models to the brain basis of vision. Computational tools in neuroscience enable us to make sense of large, heterogeneous and noisy datasets. This branch of my research brings machine-learning techniques and open source software development to neural data analysis. Specifically, I collaborate with theorists, data scientists, and experimentalists in both human neuroimaging and primate neurophysiology to study natural scene perception, visual search, motor planning, and movement sequence learning.
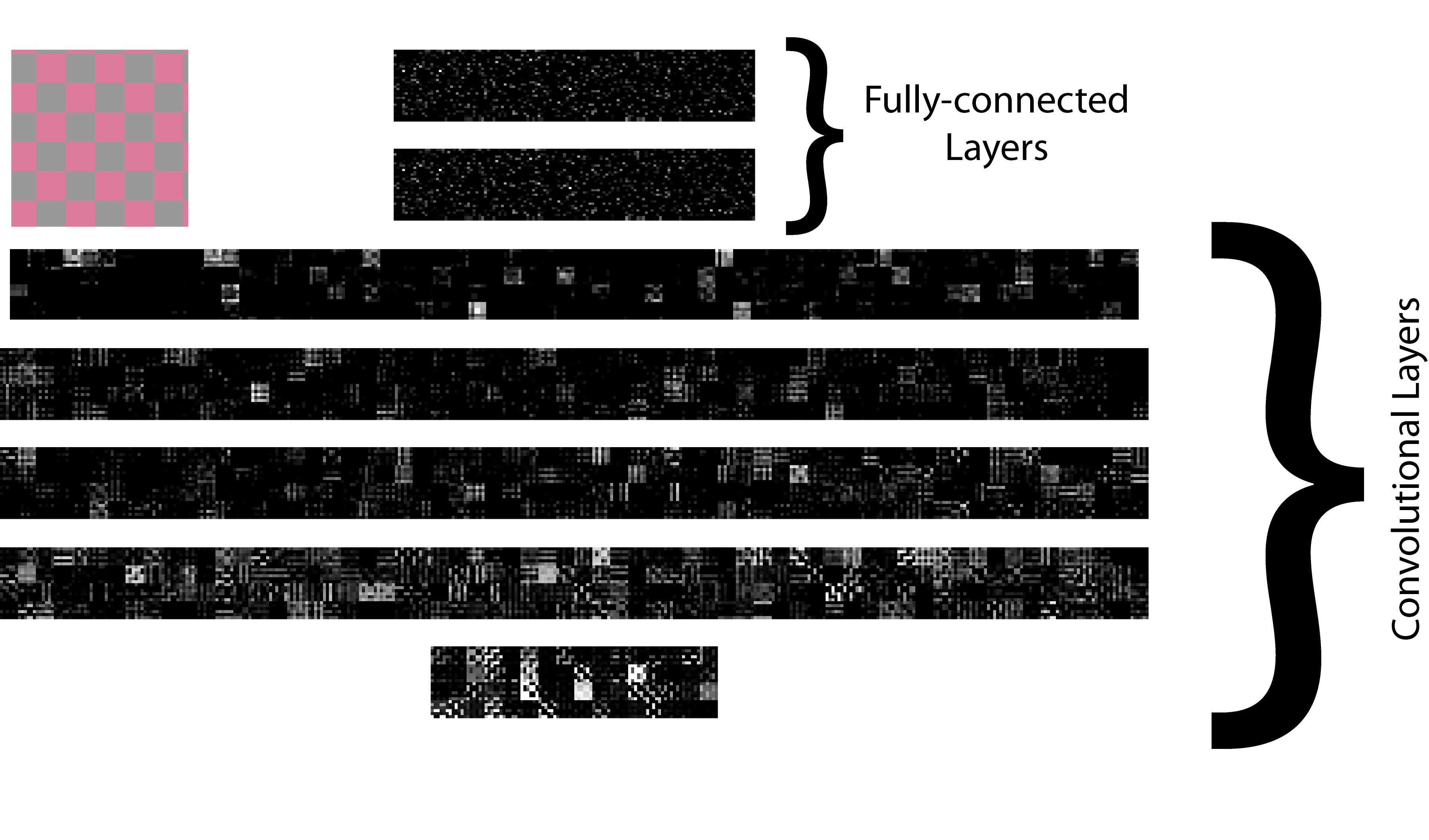
The rate of technical advancement in neuroscience will result in an avalanche of data; yet for theforeseeable future, our experiments will undersample both the animal’s behavioral repertoire and the entire variability of its brain state. This combination of data deluge and partial observability, makes the testing of even the most neurobiologically grounded theories of brain function extremely challenging. Advances in deep learning can contribute to both these problems. Modern deep neural networks have as many neurons as a larval zebrafish. They can already match human behavior in object recognition and visually guided reaching movements. Importantly, unlike animal brains, deep neural networks with complex behaviors are fully observable and controllable: we can record their state throughout learning, modify weights, dropout neurons, or rewrite their loss function. Thus, we are confronted with a choice to measure and perturb real brains imprecisely or to measure and perturb deep network models of brain-like behaviors precisely. As a sandbox for sharpening our theory, experiments and data analysis tools, my research program will integrate this approach alongside traditional computational neuroscience work to model and analyze a wide range of behaviors in visual perception and motor control.